5. Defacing Workflow¶
5.1. Introduction¶
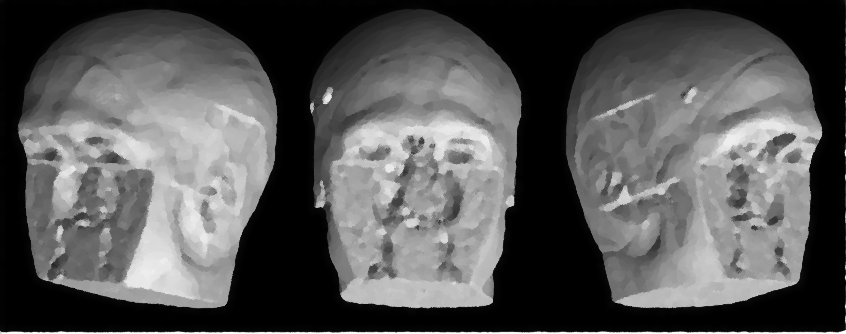
Defacing MRI data is crucial for protecting participant privacy in neuroimaging research. Structural brain MRI include facial features that can be reconstructed and used to identify individuals. Defacing removes these features while preserving brain anatomy for analysis.
5.2. Requirements¶
CPU |
RAM |
|---|---|
1 |
5 GB |
5.3. Description¶
Processing Steps
Defacing T1w image We use the UK-Biobank defacing method that is provided as part of the standard FSL distribution under the command
fsl_deface[1]. Similar to other established defacing tools, such asmri_deface[2] andpydeface[3], the method relies on linear registration to generate a mask that identifies facial voxels. Voxels within this mask are then set to zero to remove identifiable facial features. A key distinction offsl_defacecompared withmri_defaceandpydefaceis that it additionally removes the ears, providing a more comprehensive anonymization of head anatomy. This workflow is applied to T1-weighted (T1w) structural images and can be propagated to other modalities through rigid alignement.
Quality Control
Overlap score For each image, we compute the overlap ratio between the brain mask extracted using FreeSurfer’s deep‑learning–based
mri_synthstripmethod [4] and the corresponding defacing mask. Images are then sorted in ascending order of this score, allowing potential outliers to be easily identified.Manual inspection Following the overlap-based ranking, a manual quality control step is performed using the generated
defacemosaicfigure. This figure allows visual inspection of the defaced image and verification that facial/ear structures have been successfully removed. The most obvious outliers are thus removed.Thresholding The overlap score is thresholded at 5%, meaning that if the defacing mask removes a substantial portion of the brain, the preprocessing is considered invalid. Images with an overlap greater than 5% are flagged as low‑quality.
5.4. Outputs¶
The defacing directory contains subject-level results, logs, and
quality-control outputs.
The structure is organized following the brainprep ontology.
defacing/
├── dataset_description.json
├── figures
│ └── histogram_overlap.png
├── log
│ └── report_<timestamp>.rst
├── quality_check
│ └── mask_overlap.tsv
└── subjects
└── sub-01
└── ses-01
├── figures
│ └── sub-01_ses-01_run-01_mod-T1w_defacemosaic.png
├── log
│ └── report_<timestamp>.rst
├── sub-01_ses-01_run-01_mod-T1w_defacemask.nii.gz
├── sub-01_ses-01_run-01_mod-T1w_defacemask.tsv
└── sub-01_ses-01_run-01_mod-T1w_deface.nii.gz
Description of contents:
dataset_description.jsonMetadata describing the process, including versioning and processing information.figures/histogram_overlap.pngImage overlap distribution and applied threshold.logs/report_<timestamp>.rstContains group-level workflow steps and parameters.quality_check/mask_overlap.tsvTable containing the overlap score for each subject/session/run. The table includes a binaryqccolumn indicating the quality control result.subjects/sub-<id>/ses-<id>/figures/sub-01_ses-01_run-01_mod-T1w_defacemosaic.pngA visual mosaic showing defacing masks on some slices for quick quality check.subjects/sub-<id>/ses-<id>/logs/report_<timestamp>.rstContains subject-level workflow steps and parameters.subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_mod-T1w_defacemask.nii.gzThe binary mask identifying voxels removed during defacing (face and ears).subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_mod-T1w_defacemask.tsvA table containing voxel counts and physical volumes (in mm³) for the brain/defacing masks and their intersection.subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_mod-T1w_deface.nii.gzThe final defaced T1w image with facial/ear structures removed.
5.5. Featured examples¶
Defacing
Explore how to perform this analysis.