6. QuasiRAW Workflow¶
6.1. Introduction¶
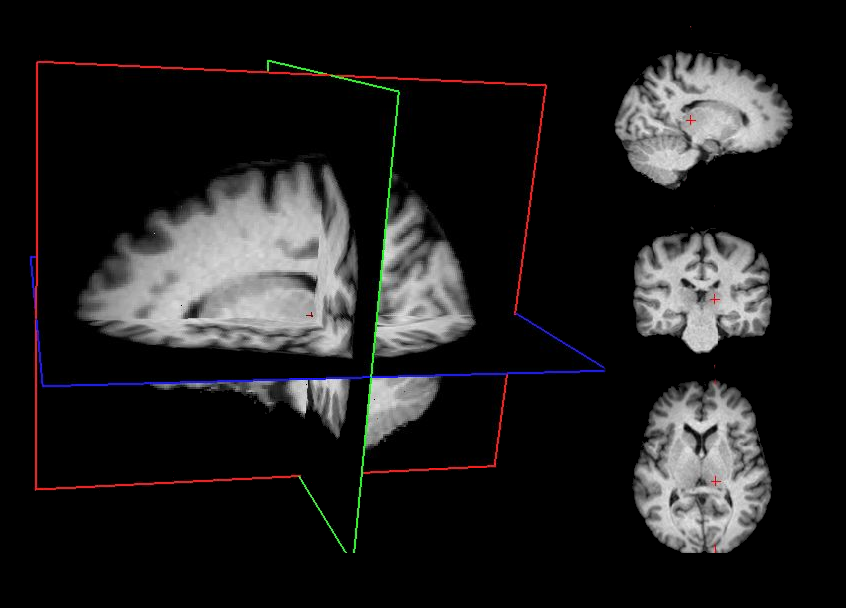
Minimally preprocessed data are generated using a standardized sequence of lightweight processing steps applied to the raw T1-weighted (T1w) MRI images. This workflow combines skull stripping, bias field correction, and spatial normalization using widely adopted neuroimaging tools. This minimal preprocessing pipeline ensures that the data are standardized and ready for subsequent processing stages while maintaining maximal fidelity to the original raw images.
6.2. Requirements¶
CPU |
RAM |
|---|---|
1 |
5 GB |
6.3. Description¶
Processing Steps
Skull stripping Brain extraction is performed using FreeSurfer’s deep learning–based
mri_synthstripmethod [1], which provides a robust and accurate removal of non-brain tissues.Bias field correction Intensity non-uniformities are corrected using the ANTs N4 algorithm [2], improving image homogeneity.
Affine registration to MNI space Spatial alignment is carried out using FSL FLIRT [3] with a 9‑degree‑of‑freedom (DOF) affine transformation (translations, rotations, and scaling; no shearing). This step registers the T1w image to the MNI template while preserving overall anatomy.
Quality Control
Correlation score For each image, we compute its correlation with the MNI template. Images are then sorted in ascending order of this score, allowing potential outliers to be easily identified.
Manual inspection Following the correlation-based ranking, generated
T1wimages at the lower end of the distribution are manually reviewed in-house. This step is performed using a PCA‑based reduction technique to detect the most obvious outliers, which are then removed.Thresholding The correlation score is thresholded at 0.5, meaning that if an image is not roughly registered to the template, the preprocessing is considered invalid. Images with a correlation lower than 0.5 are flagged as low‑quality.
6.4. Outputs¶
The quasiraw directory contains subject-level results, logs, and
quality-control outputs.
The structure is organized following the brainprep ontology.
quasiraw/
├── dataset_description.json
├── figures
│ ├── histogram_mean_correlation.png
│ └── pca.png
├── log
│ └── report_<timestamp>.rst
├── quality_check
│ ├── mean_correlations.tsv
│ └── pca.tsv
└── subjects
└── sub-01
└── ses-01
├── log
│ └── report_<timestamp>.rst
├── sub-01_ses-01_run-01_mod-T1w_affine.txt
├── sub-01_ses-01_run-01_mod-T1w_brainmask.nii.gz
└── sub-01_ses-01_run-01_T1w.nii.gz
Description of contents:
dataset_description.jsonMetadata describing the process, including versioning and processing information.figures/histogram_mean_correlation.pngImage correlation-to-template distribution and applied threshold.figures/pca.pngDisplay of the first two PCA components of the generated images.log/report_<timestamp>.rstContains group-level workflow steps and parameters.quality_check/mask_overlap.tsvTable containing the correlation score for each subject/session/run. The table includes a binaryqccolumn indicating the quality control result.quality_check/pca.tsvTable containing information on the first two PCA components.subjects/sub-<id>/ses-<id>/log/report_<timestamp>.rstContains subject-level workflow steps and parameters.subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_mod-T1w_affine.txtAffine transformation parameters (9 DOF) used to align the T1w image to the MNI template.subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_mod-T1w_brainmask.nii.gzBrain mask generated during skull stripping (e.g., via SynthStrip).subjects/sub-<id>/ses-<id>/sub-01_ses-01_run-01_T1w.nii.gzThe minimally preprocessed T1w image, including skull stripping, bias correction, and affine alignment.
6.5. Featured examples¶
Quasi RAW
Explore how to perform this analysis.